[21st Century DEVO-3] Embryonic development is the process by which a fertilized egg becomes an independent organism, an organism capable of producing functional gametes, and so a new generation. In an animal, this process generally involves substantial growth and multiple rounds of mitotic cell division; the resulting organism, a clone of the single-celled zygote, contains hundreds, thousands, millions, billions, or trillions of cells [link]. These dividing, migrating, differentiating, and sometimes dying cells that interact to form the adult and its various tissues and organ systems. The various cell types generated can be characterized by the genes that they express, the shapes they assume, the behaviors that they display, and how they interact with neighboring and distant cells (1). Based on first principles, one could imagine (at least) two general mechanisms that could lead to differences in gene expression between cells. The first would be that different cells contain different genes while the other is that while all cells contain all genes, which genes are expressed in a particular cell varies, it is regulated by molecular processes that determine when, where, and to what the levels particular genes are expressed (2). Turns out, there are examples of both processes among the animals, although the latter is much more common.
The process of discarding genomic DNA in somatic cells is known as chromatin diminution. During the development of the soma, but not the germ line, regions of the genome are lost. In the germ line, for hopefully obvious reasons, the full genome is retained. The end result is that somatic cells contain different subsets of genes and non-coding DNA compared to the full genome. The classic case of chromosome diminution was described in the parasitic nematode of horses, now named Parascaris univalens (originally Ascaris megalocephala) by Theodore Boveri in 1887 (reviewed in Streit and Davis, 2016)[pdf link]. Based on its occurrence in a range of distinct animal lineages, chromatin diminution appears to be an emergent rather than an ancestral trait, that is, a trait present in the common ancestor of the animals.
While, as expected for an emergent trait, the particular mechanism of chromatin diminution appears to vary between different organisms: the best characterized example occurs in Parascaris. In the somatic cell lineages in which chromatin diminution occurs, double-stranded breaks are made in chromosomal DNA molecules, and teleomeric sequences are added to ends of the resulting DNA molecules (↓).

You may have learned that chromosomes interact with spindle microtubules through a localized regions on the chromosomes, known as centromeres. Centromeres are identified through their association with proteins that form the kinetochore, which is a structure that mediates interactions between condensed chromosomes and mitotic (and meiotic) spindle microtubules. While many organisms have a discrete spot-like (localized) centromere, in many nematodes centromere-binding proteins are found distributed along the length of the chromosomes, a situation known as a holocentric centromere. At higher resolution it appears that centromere components are preferentially associated with euchromatic, that is, molecularly accessible chromosomal regions, which are (typically) the regions where most expressed genes are located. Centromere components are largely excluded from heterochromatic (condensed and molecularly inaccessible) chromosomal regions. After chromosome fragmentation, those DNA fragments associated with centromere components can interact with the spindle microtubules and are accurately segregated to daughter cells during mitosis, while those, primarily heterochromatic fragments (without associated centromeric components) are degraded and lost. In contrast the integrity of the genome is maintained in those cells that come to form the germ line, the cells that can undergo meiosis to produce gametes. Looking forward to the reprogramming of somatic cells (the process of producing what are known as induced pluripotent stem cells – iPSCs), one prediction is that it should not be possible to reprogram a somatic cell that has undergone chromatin diminution to form a functional germ line cell – you should be able to explain why, or what would have to be the case for such reprogramming to be successful.
The origins of cellular asymmetries: Clearly, there must be differences between the cells that undergo chromatin diminution and those that do not; at the very least the nuclease(s) that cuts the DNA during chromatin diminution will need to be active in somatic cells and inactive in germ line cells, or it may simply not be present – the genes that encode it are not expressed in germ line cells. We can presume that similar cytoplasmic differences play a role in the differential regulation of gene expression in different cell types during the development of organisms in which the genome remains intact in somatic cells. So how might such asymmetries arise? There are three potential, but certainly not mutually exclusive, mechanisms that can lead to cellular/cytoplasmic asymmetries: they can be inherited based on pre-existing asymmetries in the parental cell, they could emerge based on asymmetries in the signaling environments occupied by the two daughters, or they could arise from stochastic fluctuations in gene expression (see Chen et al., 2016; Neumüller and Knoblich, 2009).

One example of how an asymmetry can be established occurs in the free-living nematode Caenorhabditis elegans, where the site of sperm fusion with the egg leads to the recruitment and assembly of proteins around the site of sperm entry, the future posterior side of the embryo. After male and female pronuclei fuse, mitosis begins and cytokinesis divides the zygote into two cells; the asymmetry initiated by sperm entry leads to an asymmetric division (↑); the anterior AB blastomere is larger, and molecularly distinct from the smaller posterior P1 blastomere. These differences set off a regulatory cascade, in which the genes expressed at one stage influence those expressed subsequently, and so influence subsequent cell divisions / cell fate decisions.
Other organisms use different mechanisms to generate cellular asymmetries. In organisms that have external fertilization, such as the clawed frog Xenopus, development proceeds rapidly once fertilization occurs. The egg is large, since in contains all of the materials necessary for the formation until the time that the embryo can feed itself. The early embryo is immotile and vulnerable to predation, so early development in such species tends to be rapid, and based on materials supplied by the mother (leading to maternal effects on subsequent development). In such cases, the initial asymmetry is built into the organization of the oocyte.
Formed through a mitotic division the primary oocyte enters meiotic prophase I, during which it undergoes a period of growth. Maternal and paternal chromosomes align (syngamy) and undergo crossing-over (recombination). The oocyte contains a single centrosome, a cytoplasmic structure that surrounds the centrioles of the oocyte’s inherited mitotic spindle pole. Cytoplasmic components become organized around the pole and then move from the pole toward the cell cortex (↓ image from Gard and Klymkowsky, 1998); this movement defines an “animal-vegetal” axisof the oocyte, which upon fertilization will play a role in generating the head-tail (anterior-posterior) and back-belly (dorsal-ventral) axes of the embryo and adult.
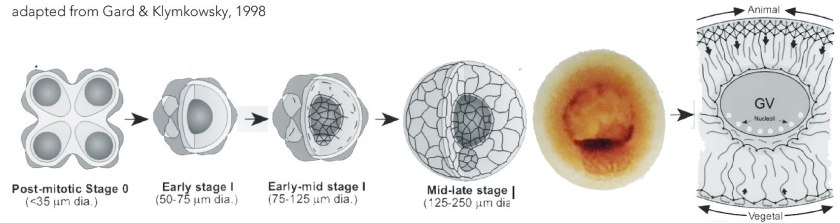
The primary oocyte remains in prophase I throughout oogenesis. The asymmetry of the oocyte becomes visible through the development of a pigmented animal hemisphere, largely non-pigmented vegetal hemisphere, and an large (~300 um diameter) and off-centered nucleus (known as the germinal vesicle or GV)(3). Messenger RNA molecules, encoding different polypeptides, are differentially localized to the animal and vegetal regions of the late stage oocyte. The translation of these mRNAs is regulated by factors activated by subsequent developmental events, leading to molecular asymmetries between embryonic cells derived from the animal and vegetal regions of the oocyte. In preparation for fertilization, the oocyte resumes active meiosis, leading to the formation of two polar bodies and the secondary oocyte, the egg. Fertilization occurs within the pigmented animal hemisphere; the site of sperm entry (↓) produces a second driver of asymmetry, in addition to the animal-vegetal axis, albeit through a mechanism distinct from that used in C. elegans (De Domenico et al., 2015).

Asymmetries in oocytes and eggs, and sperm entry points are not always the primary drivers of subsequent embryonic differentiation. In the mouse, and other placental mammals, including humans, embryonic development occurs within, and is supported by and dependent upon the mother. The mouse (mammalian) egg appears grossly symmetric, and sperm entry itself does not appear to impose an asymmetry. Rather, as the zygote divides, the first cells formed appear to be similar to one another. As cell division continue, however, some cells find themselves on the surface while others are located within the interior of the forming ball of cells, or morula (↓).

These two cell populations are exposed to different environments, environments that influence patterns of gene expression. The cells on the surface differentiate to form the trophectoderm, which in turn differentiates into extra-embryonic placental tissues, the interface between mother and developing embryo. The internal cells becomes the inner cell mass, which differentiate to form the embryo proper, the future mouse (or human). Early on inner cell mass cells appear similar to one another, but they also experience different environments, leading to emerging asymmetries associated with the activation of different signaling systems, the expression of different sets of genes, and difference in behavior – they begin the process of differentiating into distinct cell lineages and types forming, as embryogenesis continues, different tissues and organs.
The response of a particular cell to a particular environment will depend upon the signaling molecules present, typically expressed by neighboring cells, the signaling molecule receptors expressed by the cell itself, and how the binding of signaling molecules to receptors alters receptor activity or stability. For example, an activated receptor can activate (or inhibit) a transcription factor protein that could influence the expression of a subset of genes. These genes may themselves encode regulators of transcription, signals, signal receptors, or modifiers of the cellular localization, stability, activity, or interactions with other molecules. While some effects of signal-receptor interactions can be transient, leading to reversible changes in cell state (and gene expression), during embryonic development activating and responding to a signal generally starts a cascade of effects that leads to irreversible changes, and the formation of altered differentiated states.
A cell’s response to a signal can be variable, and influenced by the totality of the signals it receives and its past history. For example, a signal could lead to a decrease in the level of a receptor, or an increase in an inhibitory protein, making the cell unresponsive to the signal (a negative feedback effect) or more sensitive (a positive feedback effect) or could lead to a change in its response to a signal – different genes could be regulated as time goes by following the signal. Such emerging patterns of gene expression, based on signaling inputs, are the primary driver of embryonic development.
footnotes:
- Not all genes are differentially expression, however – some genes, known as housekeeping genes, are expressed in essential all cells.
- Hopefully it is clear what the term “expressed” means – namely that part of the gene is used to direct the synthesis of RNA (through the process of transcription (DNA-dependent, RNA polymerization). Some such RNAs (messenger or mRNAs) are used to direct the synthesis of a polypeptide through the process of translation (RNA-directed, amino acid polymerization) others do not encode polypeptides, such non-coding RNAs (ncRNAs) can play roles in a number of processes, from catalysis to the regulation of transcription, RNA stability, and translation.
- Eggs are laid in water and are exposed to the sun; the pigmentation of the animal hemisphere is thought to protect the oocyte/zygote/early embryo’s DNA from photo-damage.
Literature cited
Chen et al., (2016). The ins (ide) and outs (ide) of asymmetric stem cell division. Current opinion in cell biology 43, 1-6.
De Domenico et al., (2015). Molecular asymmetry in the 8-cell stage Xenopus tropicalis embryo described by single blastomere transcript sequencing. Developmental biology 408, 252-268.
Gard & Klymkowsky. (1998). Intermediate filament organization during oogenesis and early development in the clawed frog, Xenopus laevis. In Intermediate filaments (ed. H. Herrmann & J. R. Harris), pp. 35-69. New York: Plenum.
Neumüller & Knoblich. (2009). Dividing cellular asymmetry: asymmetric cell division and its implications for stem cells and cancer. Genes & development 23, 2675-2699.
Streit & Davis. (2016). Chromatin Diminution. In eLS: John Wiley & Sons Ltd, Chichester.


4 thoughts on “Establishing Cellular Asymmetries: a biofundamentalist perspective”